DiffDock-L
Dock a ligand onto any protein receptor with high accuracy.
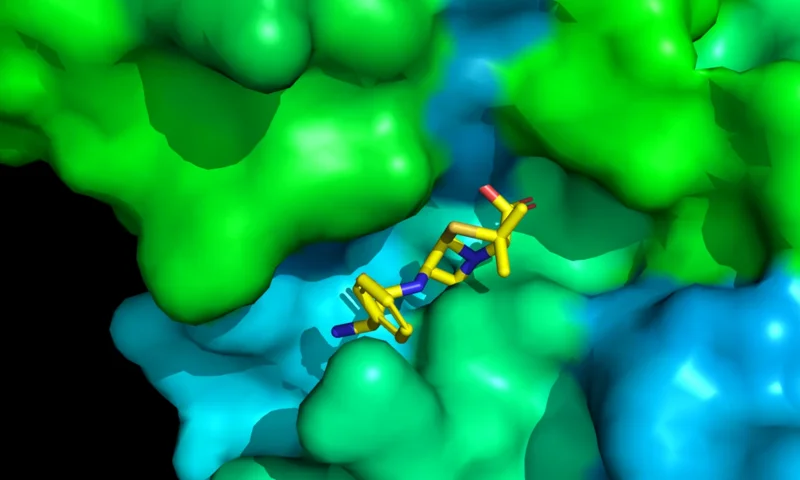
DiffDock-L is the newer and better version of DiffDock, a state of the art method for molecular docking and drug binding structure prediction. This service uses DiffDock-L and AutoDock VINA to predict protein-ligand interactions with high accuracy.
Estimated resource cost: 0.000527
Categories: Ligand/Drug Design & Screening, Molecular Docking & Interactions
Tags: Affinity Prediction Protein–Ligand Docking Virtual Screening
Key Capabilities
- Utilizes the improved DiffDock-L implementation.
- Predict ligand binding to a protein target of interest.
- Can be used for drug design as well as domain identification.
- Includes an interactive 3D protein viewer with the docked ligand.
Runtime Statistics
| Metric | Value |
|---|---|
| runtime_mean | 469 |
| runtime_median | 388 |
| runtime_std | 374 |
| runtime_90th_percentile | 855 |
| runtime_max | 13755 |
Similar Tools
- AutoDock Vina (smina)
- DynamicBind
- GNINA
- SPRINT

Ready to submit your job?
Review your configuration, then confirm the estimated credit cost before you run the job. Note that credit estimates are not guaranteed and runtime can vary depending on inputs and settings.
Estimated Credits: 0.000527
Invite-only, limited-time access. Please contact ztang@getantibody.com.
Invite-only, limited-time access. Please contact ztang@getantibody.com.