PocketFlow
PocketFlow is a Deep Generative Model that generates ligands for target protein binding pockets.
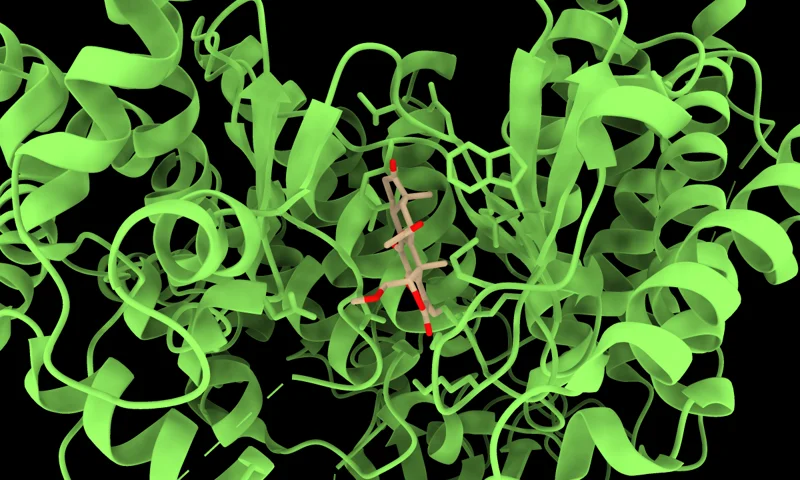
PocketFlow generates ligands for target protein binding pockets. PocketFlow considers chemical knowledge from the CrossedDock and ZINC 3D datasets during the molecular generation process.
Estimated resource cost: 0.0002945
Categories: Ligand/Drug Design & Screening
Tags: Small Molecules Virtual Screening
Key Capabilities
- PocketFlow can generate hundreds of small molecules in minutes.
- Requires only a target protein binding pocket in PDB file format.
- Returns ligands in SMI and SDF file format along with relevant metrics and graphs.
Runtime Statistics
| Metric | Value |
|---|---|
| runtime_mean | 735 |
| runtime_median | 404 |
| runtime_std | 1016 |
| runtime_90th_percentile | 2009 |
| runtime_max | 11559 |
Similar Tools
- GenMol
- QEPPI
- SPRINT
- AutoDock Vina (smina)

Ready to submit your job?
Review your configuration, then confirm the estimated credit cost before you run the job. Note that credit estimates are not guaranteed and runtime can vary depending on inputs and settings.
Estimated Credits: 0.0002945
Invite-only, limited-time access. Please contact ztang@getantibody.com.
Invite-only, limited-time access. Please contact ztang@getantibody.com.